
EPPO Workshop on setting Ct cut-off values for real-time PCR
Paris, 2013-11-11/12
Background
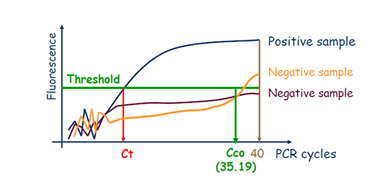 There was a recommendation in the instructions to authors of EPPO Diagnostic Protocols that for real-time PCR test descriptions, a Ct cut-off value should be provided (although a comment should always be included in this test description that ‘the cycle cut-off value needs to be verified in each laboratory when implementing the test’). The initial aim of the Workshop was consequently to prepare recommendations to harmonize the determination of Ct cut-off values.
There was a recommendation in the instructions to authors of EPPO Diagnostic Protocols that for real-time PCR test descriptions, a Ct cut-off value should be provided (although a comment should always be included in this test description that ‘the cycle cut-off value needs to be verified in each laboratory when implementing the test’). The initial aim of the Workshop was consequently to prepare recommendations to harmonize the determination of Ct cut-off values.
Main conclusions
Eight presentations from experts from six countries were made on the experience with real-time PCR in different laboratories (including presentations from the animal health and GMO fields). In the presentations, experts presented specific situations where they considered Ct cut-off values were necessary. However, it became clear from these presentations that setting a Ct cut-off value is not necessary in most situations and that the presence of an exponential curve is usually an indication of the presence of the target for tests which are specific.
Consequently it was recommended that the requirement made in the instructions to authors to specify a Ct cut-off value for all real-time PCR tests should be amended. A proposal was agreed during the meeting. This proposal was presented to the Panel on Diagnostics and Quality Assurance (2013-11-13/15) and was agreed. The modifications have been included in the revised instructions to authors of EPPO Diagnostic Protocols (Appendix 2 point 3.2).
It was recognized that in certain cases use of Ct cut-off may be recommended by the authors of the test description (e.g. when it is known that healthy material gives late exponential curves or there may be issues with specificity). However, reasons for exponential curves appearing in late cycles (late signals) should be investigated (e.g. by sequencing, real-time PCR targeting another region of the genome, or any other method with sufficient analytical sensitivity) and not automatically considered as negative samples.
Data (including data on the occurrence of late signals and their confirmation/refutation) should be collected by laboratories to allow continuous improvement of the test and of its interpretation. This may require the adjustment of the previously determined Ct cut-off value. As stated in the redrafted instructions to authors it is essential to recall that Ct cut-off value is equipment, material and chemistry dependent consequently it needs to be verified in each laboratory when implementing the test.
Participants underlined that the analytical sensitivity (limit of detection) value is needed to implement a test in a laboratory and should be stated, preferably with information on how it was determined. The Workshop agreed that this limit of detection should preferably be expressed in biologically relevant terms (e.g. number of specimens, cells/mL, relative infection rates), however it can also be expressed as amount of target DNA/RNA (ng or copies).
Finally it was recalled that a laboratory should perform a verification of any new test introduced in their testing programme (see PM 7/098 Specific requirements for laboratories preparing accreditation for a plant pest diagnostic activity). This verification includes the requirement of testing 8 samples at the limit of detection (for viruses, viroids and phytoplasmas, this should be at low level). It was also recalled that appropriate controls should be included in each series of nucleic acid extraction and amplification and give expected results.
The Workshop recommended that a specific session is organized on validation and verification of real-time PCR tests at the EPPO Workshop on Quality Assurance (York, UK, 2014-02-18/20). Speakers agreed to contribute to an article to be published in the EPPO Bulletin to share their experience with a wider audience.
The importance of access to reference material for development, validation and verification of tests and preparation of appropriate controls was underlined. The Workshop noted that an EU project on reference collections (Q-collect) started on the 1st of October 2013. It was strongly suggested that the issue of availability of reference material (including samples with known quantities of non-culturable organisms, target DNA inserted in plasmids) is addressed during this project.
Presentations
Determination of LOD and cycle cut-off in real-time PCR detecting culturable and non-culturable target organisms
Ms Dreo T. (SI)
Real-time PCR in animal health diagnosis according to XP U47-600 Standard: no Ct cut-off value but LOD in number of copies of target sequences
Ms Pelletier C. (FR)
Setting Ct cut-off values using Test Performance Study data
Mr Van De Vossenberg B. (NL)
A method to determine cycle cut-off in real time PCR for regulated plant pathogens
Ms Chandelier A. (BE)
Determination of a cut-off value based on the use of a limit of detection positive control
Mr Ioos R. (FR)
Report on the experiences gathered from establishing real-time PCR assays to detect Dickeya spp. (detection of DNA) and potato viruses (detection of RNA)
Mr Pastrik K.H. (DE)
Ct values: method validation and decision making
Ms Weekes R.(GB)
The approach to setting Ct cut-off values used in real-time PCR testing for GMOs was also presented by Ms Anthoine (FR).
